📝 Paper Summary
Protein Language Models (PLMs)
Mechanistic Interpretability
Efficient Inference
The paper demonstrates that protein language models rely more on semantic than positional information compared to natural language models and leverages this via an early-exit strategy to improve both accuracy and efficiency.
Core Problem
Protein language models (PLMs) are often treated identically to natural language models (NLMs) despite fundamental differences in their data domains (e.g., sequence length, vocabulary size), leading to suboptimal utilization of their internal representations.
Why it matters:
- Proteins have a small vocabulary (20 amino acids) but rich functional spaces, unlike natural language's large vocabulary and human-defined semantics
- Standard inference uses only the final layer, but intermediate layers in PLMs often contain richer biological information that is currently wasted
- Blindly applying NLP architectures without understanding domain-specific mechanistic differences limits the potential for biologically grounded model improvements
Concrete Example:
In natural language, early-exit strategies usually trade accuracy for speed. However, this paper shows that for protein tasks like non-structural property prediction, exiting at intermediate layers actually *increases* accuracy (e.g., +7.01% on a specific task) because the final layers may over-process or dilute critical biological signals found earlier.
Key Novelty
Domain-Specific Attention Analysis & PLM Early-Exit
- Directly compares attention mechanisms in PLMs and NLMs by decomposing attention into positional and semantic components, revealing that PLMs prioritize semantic content (amino acid identity/context) over position
- Adapts early-exit inference—typically a speed-accuracy trade-off in NLP—to PLMs, demonstrating it acts as a performance *booster* for protein tasks by retrieving better representations from middle layers
Architecture
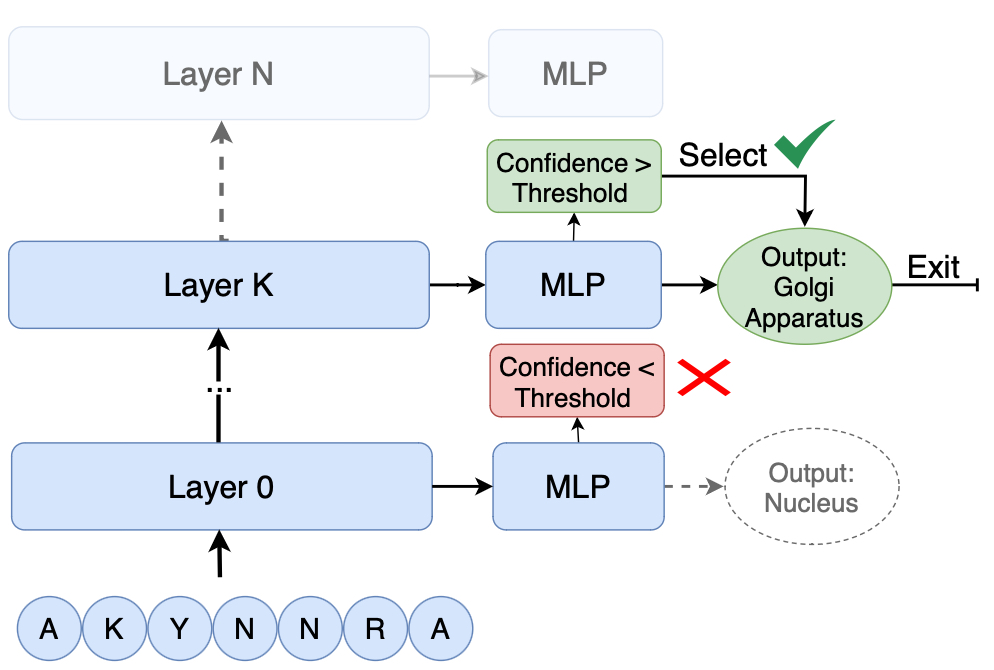
Schematic of the Early-Exit mechanism applied to a Protein Language Model.
Evaluation Highlights
- Achieved performance gains ranging from 0.4 to 7.01 percentage points across various non-structural protein property prediction tasks using early-exit
- Improved computational efficiency by over 10% across models while simultaneously increasing accuracy
- Revealed distinct attention patterns: PLMs exhibit higher semantic-to-positional attention ratios than their NLM counterparts, indicating different information processing mechanisms
Breakthrough Assessment
7/10
Provides a valuable mechanistic insight into how PLMs differ from NLMs and successfully turns a standard efficiency technique (early-exit) into a performance-enhancing tool for biology, though the method itself is an adaptation.