📝 Paper Summary
Clinical Information Extraction
Low-resource NLP (German Medical Text)
Few-shot Learning
For German clinical section classification with minimal data, a general-domain PLM adapted to the medical domain via further-pretraining and prompted with PET significantly outperforms traditional sequence classifiers.
Core Problem
Extracting information from unstructured clinical text (doctor's letters) is difficult due to high annotation costs, strict privacy regulations preventing external model usage, and limited compute resources in hospitals.
Why it matters:
- Clinical data is highly sensitive and must be processed on-premise, often barring the use of powerful API-based LLMs like GPT-4
- German clinical NLP is a lower-resource domain compared to English, lacking the massive annotated datasets required for standard supervised learning
- Medical professionals require transparent, interpretable model decisions, which black-box deep learning models often fail to provide
Concrete Example:
A standard sequence classifier trained on only 20 examples fails to distinguish 'Anamnese' (history) from 'Zusammenfassung' (summary) because they share tokens like 'patient' and 'admission'. The proposed prompt-based approach with domain adaptation correctly classifies these by leveraging contextual patterns and structural knowledge.
Key Novelty
Domain-Adapted Prompting with PET (Pattern-Exploiting Training)
- Combine lightweight Prompt-Based Learning (PET) with Further-Pretraining on domain-specific clinical text to maximize performance from minimal labeled examples (few-shot)
- Demonstrate that starting with a general-language model and adapting it to the clinical domain works better than starting with a specialized medical model pretrained from scratch on limited data
- Use Shapley values not just for explanation, but to identify and correct training data biases (e.g., specific tokens acting as false shortcuts)
Architecture
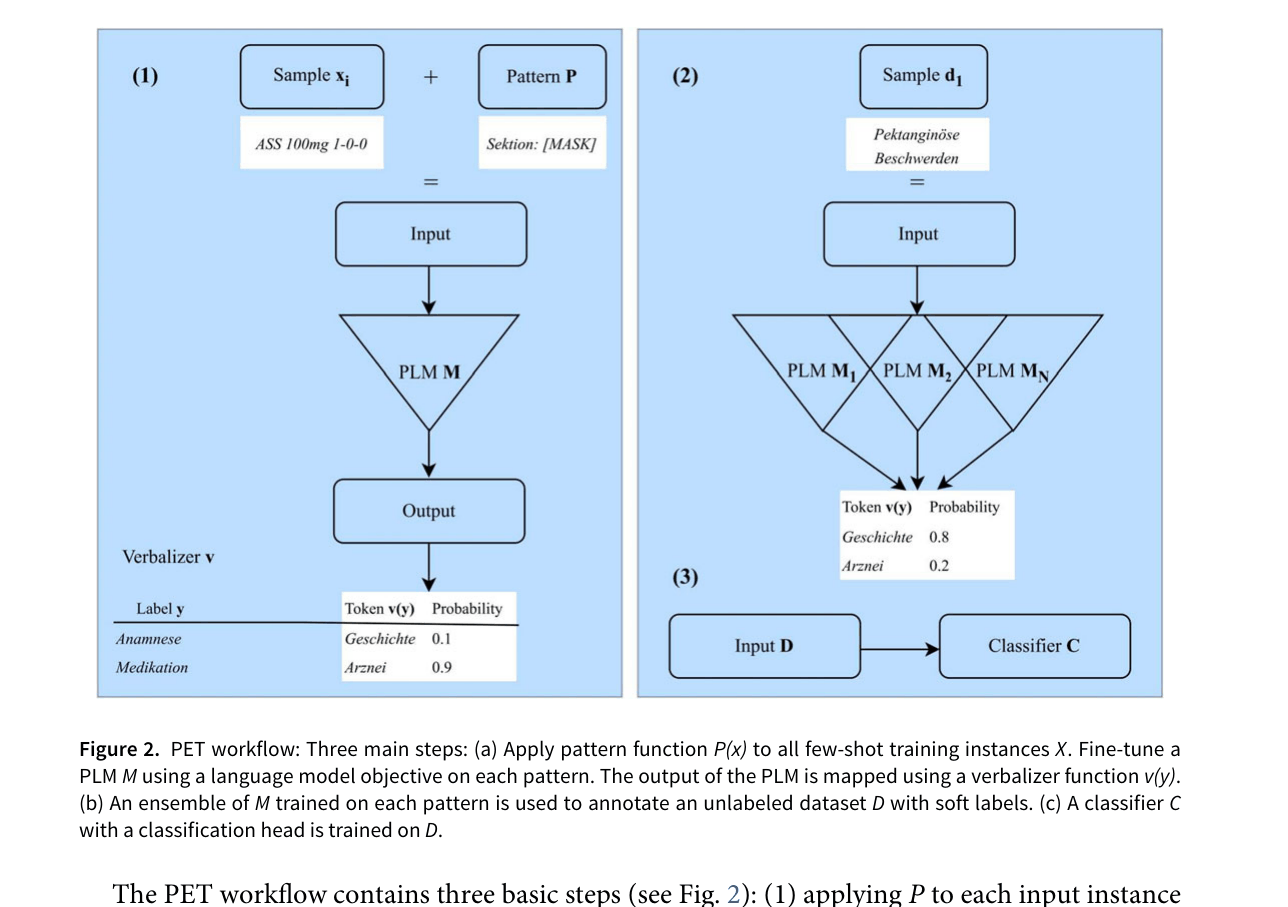
The PET (Pattern-Exploiting Training) workflow applied to clinical text
Evaluation Highlights
- With only 20 training shots per class, the proposed PET approach achieves 79.1% accuracy, outperforming a traditional sequence classifier (48.6%) by +30.5 percentage points
- Further-pretraining a general German model (gbert) on clinical data yields better few-shot performance than using a model pretrained on medical data from scratch (medbertde)
- Using Shapley values for model selection and optimization further boosts accuracy to 84.3% in the 20-shot setting
Breakthrough Assessment
7/10
Strong pragmatic contribution for low-resource clinical NLP. Demonstrates that adapting general models is superior to specialized small models for German, and effectively integrates interpretability for model improvement.