📝 Paper Summary
Reinforcement Learning for Reasoning
Model Calibration
AI for Science
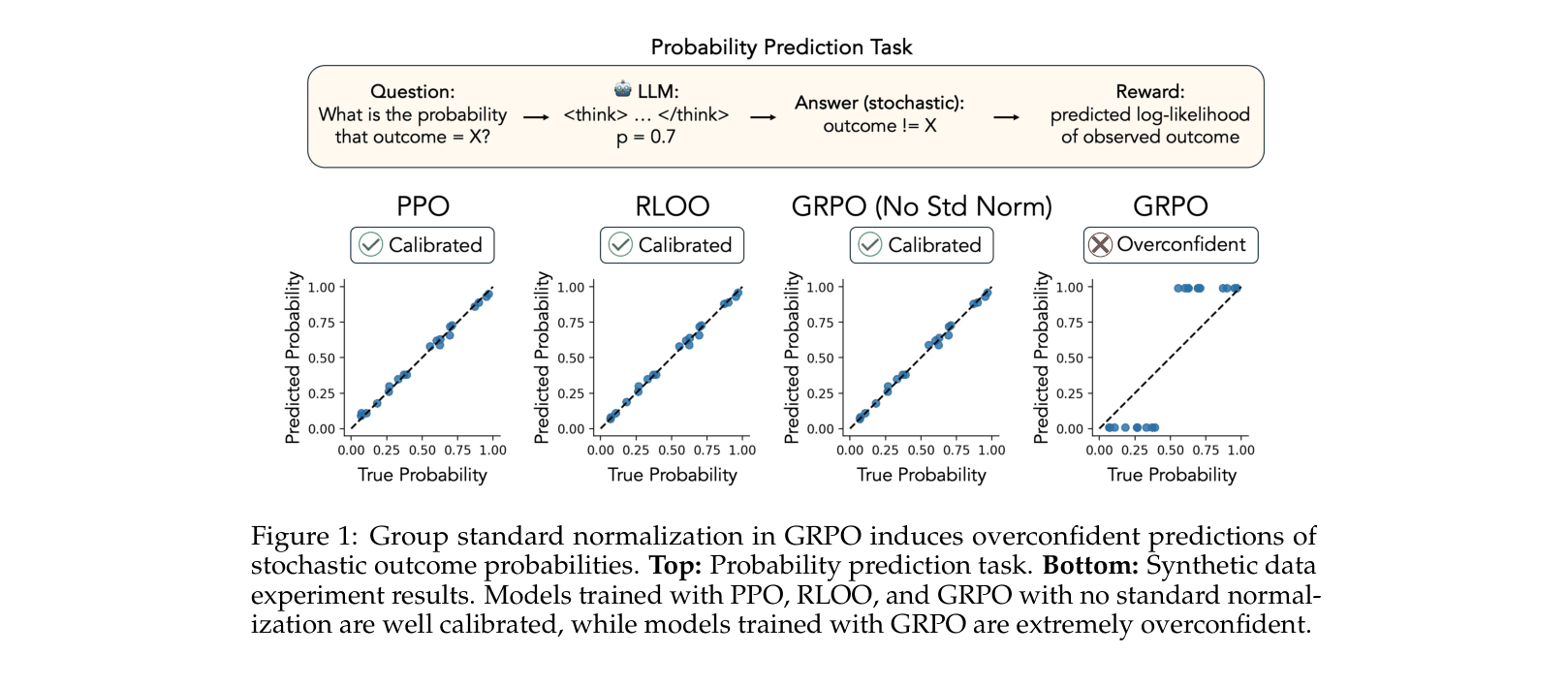
Group Relative Policy Optimization (GRPO) inherently causes language models to become overconfident in stochastic domains due to biased advantage normalization, whereas PPO and RLOO remain calibrated.
Core Problem
Reinforcement learning methods like GRPO excel in deterministic domains (e.g., math) but fail in stochastic settings (e.g., scientific experiments), inducing extreme overconfidence in predicted probabilities.
Why it matters:
- Scientific reasoning requires models to accurately estimate uncertainty and probabilities, not just provide binary answers
- Standard RL reasoning methods effectively break model calibration, making them unreliable for high-stakes decision-making or hypothesis generation
- Current trends in 'reasoning' models assume verifiable deterministic ground truth, leaving a gap for probabilistic real-world tasks
Concrete Example:
In a CRISPR experiment where a gene perturbation has a 70% chance of an effect, a GRPO-trained model is driven to predict near 100% or 0%, while PPO correctly converges to the 70% probability.
Key Novelty
Bias Identification in GRPO Normalization
- Identifies that the standard normalization term in GRPO's advantage estimator (dividing by group standard deviation) creates a policy-dependent bias
- Demonstrates that this bias creates a feedback loop: as the policy concentrates, the normalization term amplifies the reward signal for overconfident predictions
- Proposes removing group standard normalization from GRPO to restore unbiasedness and achieve calibration comparable to PPO and RLOO
Architecture
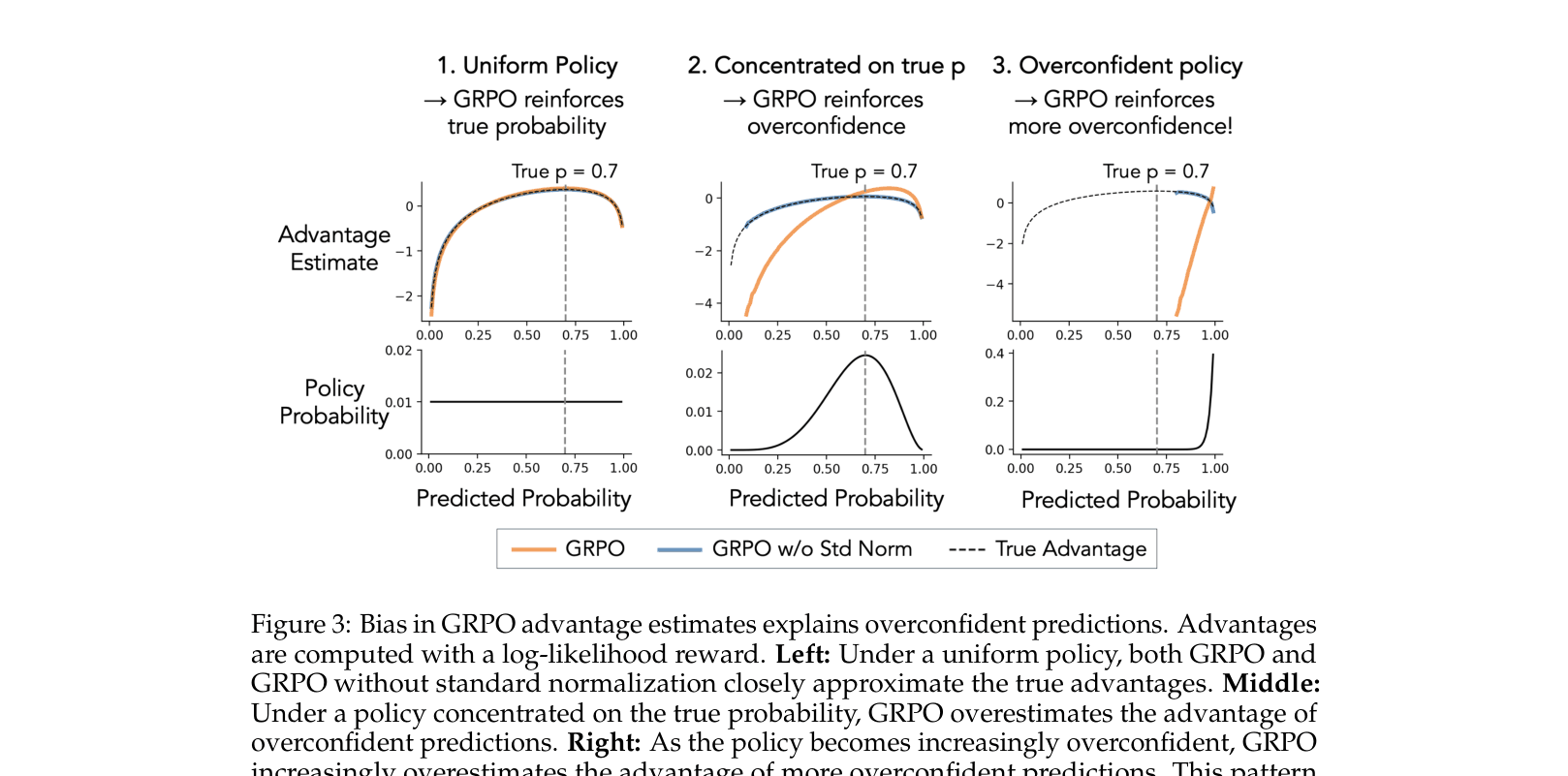
Analysis of bias in GRPO advantage estimates compared to True Advantage and No-Standardization GRPO
Evaluation Highlights
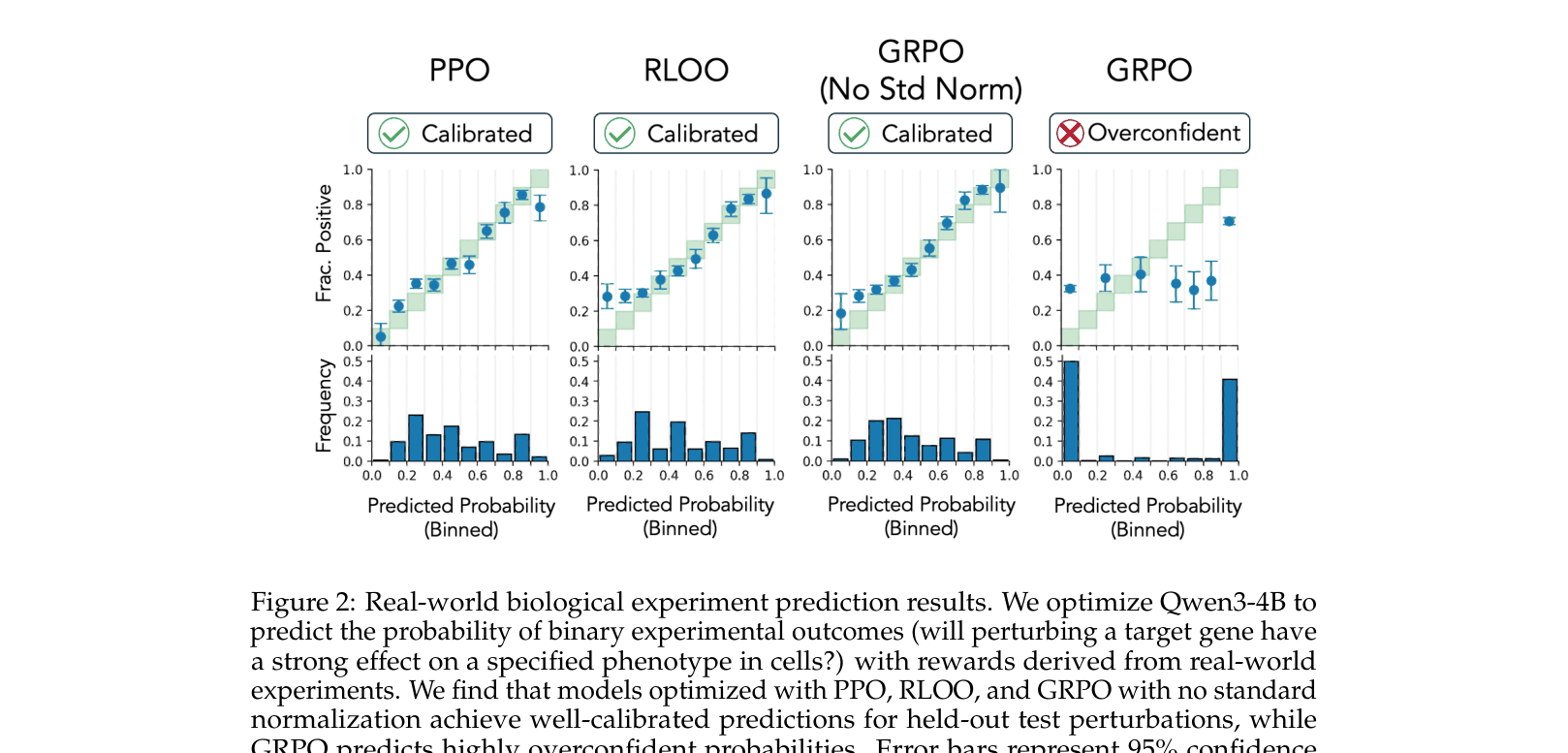
- GRPO reduces Expected Calibration Error (ECE) from 0.292 (standard) to 0.036 (no normalization) on real-world CRISPR tasks using Qwen3-4B
- Standard GRPO yields an AUROC of 0.69 on CRISPR data, significantly worse than PPO (0.72) and RLOO (0.72)
- On synthetic data, GRPO produces extreme overconfidence (ECE 0.239) while PPO and RLOO remain perfectly calibrated (ECE < 0.005)
Breakthrough Assessment
7/10
Provides a crucial diagnostic and fix for a popular algorithm (GRPO) in a new domain (stochastic reasoning). Theoretical analysis is sound and experiments are clear, though scope is limited to calibration.