📝 Paper Summary
Machine Learning for Molecular Dynamics
Neural Potentials / Force Fields
Graph Neural Networks (GNNs)
A masked pretraining method for GNNs that learns molecular structure by recovering the spatial information of masked-out atoms, improving downstream force and energy prediction accuracy.
Core Problem
Training accurate neural potentials requires expensive DFT data, and existing pretraining methods like denoising struggle with scaling or selecting appropriate noise levels for complex systems like water.
Why it matters:
- Ab initio methods (DFT) are computationally prohibitive for large systems, necessitating efficient machine learning surrogates.
- Collecting large DFT datasets is expensive; improving data efficiency through pretraining allows high accuracy with fewer labels.
- Existing pretraining methods (e.g., denoising) can be unstable or ineffective on systems with complex intra- and inter-molecular interactions like water.
Concrete Example:
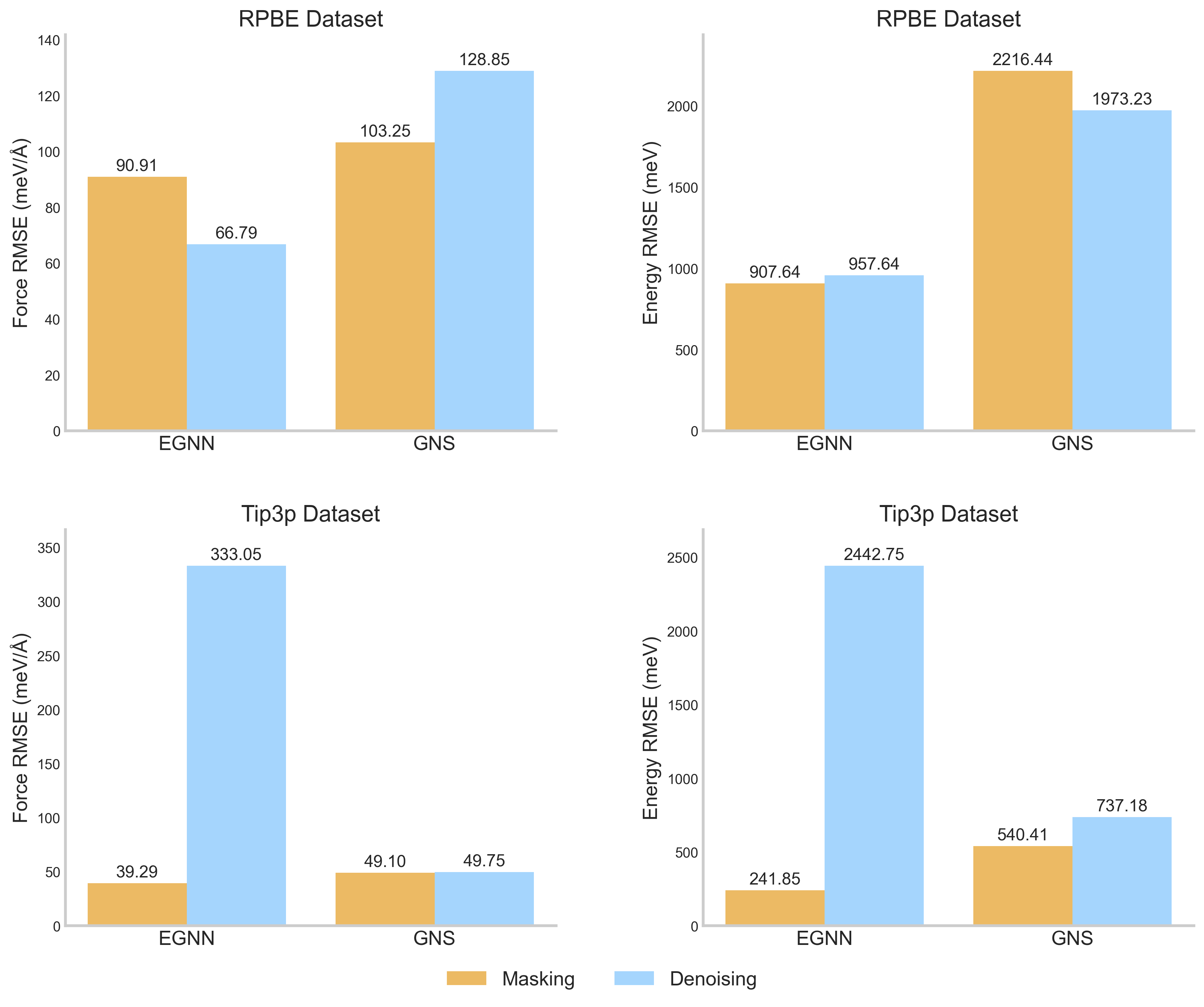
In water systems, denoising pretraining often fails to scale: for the EGNN model on the Tip3p dataset, denoising leads to poor convergence (RMSE rising to ~2442 meV/Å), whereas the proposed masking strategy remains stable and accurate (RMSE ~241 meV).
Key Novelty
Hydrogen Atom Masking for 3D Molecular Graphs
- Selectively masks one Hydrogen atom from water molecules and tasks the GNN with predicting its spatial displacement relative to the rest of the molecule.
- Forces the network to learn inherent structural and physical priors (like bond lengths and angles) by reconstructing missing geometry, rather than just denoising coordinates.
- Uses negative Cosine Similarity as the loss function, focusing on relative positioning (direction) rather than absolute distance, making it robust to scale variations.
Architecture
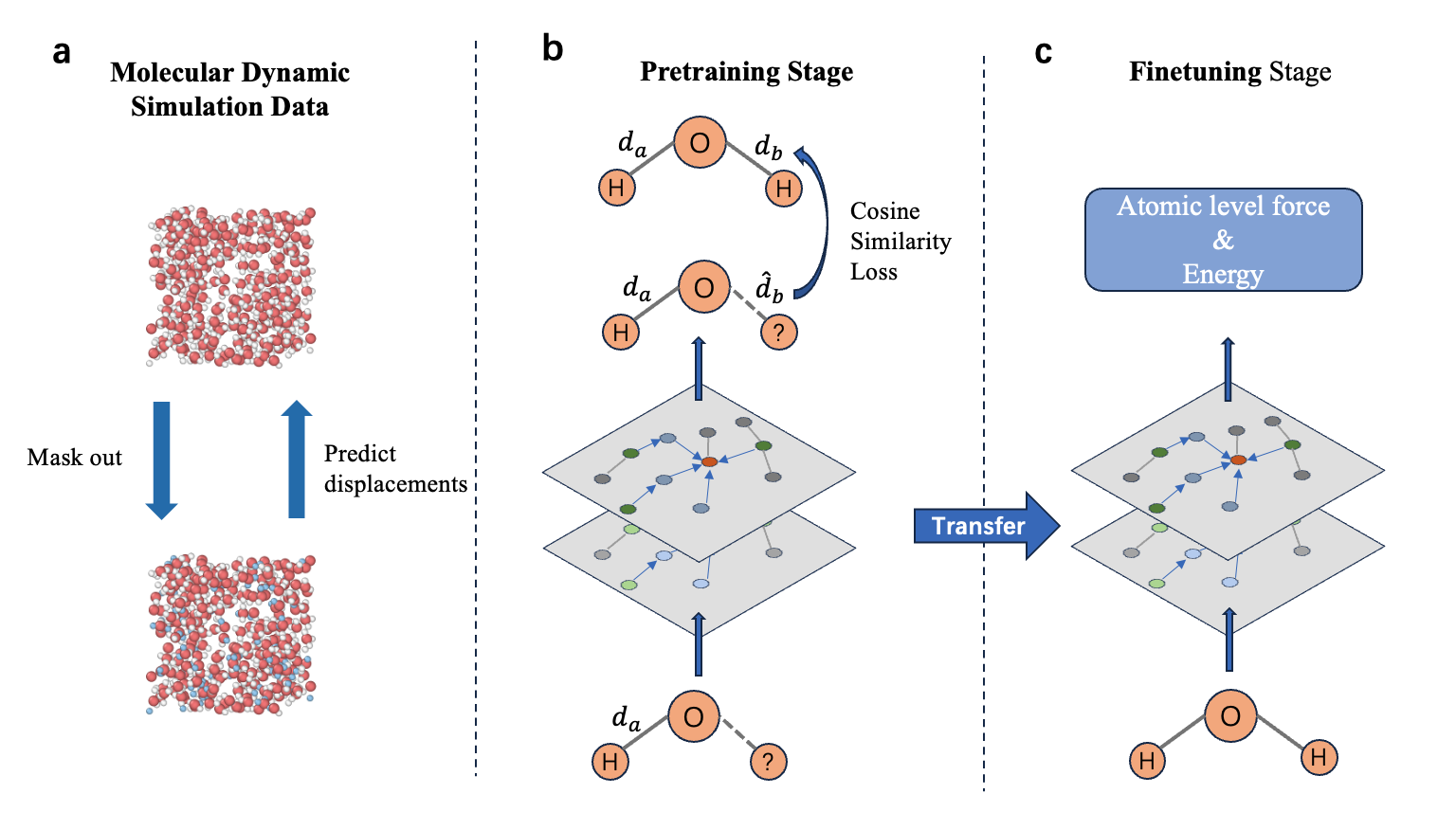
Illustration of the two-stage training pipeline: Pretraining via Hydrogen masking and Finetuning on forces/energy.
Evaluation Highlights
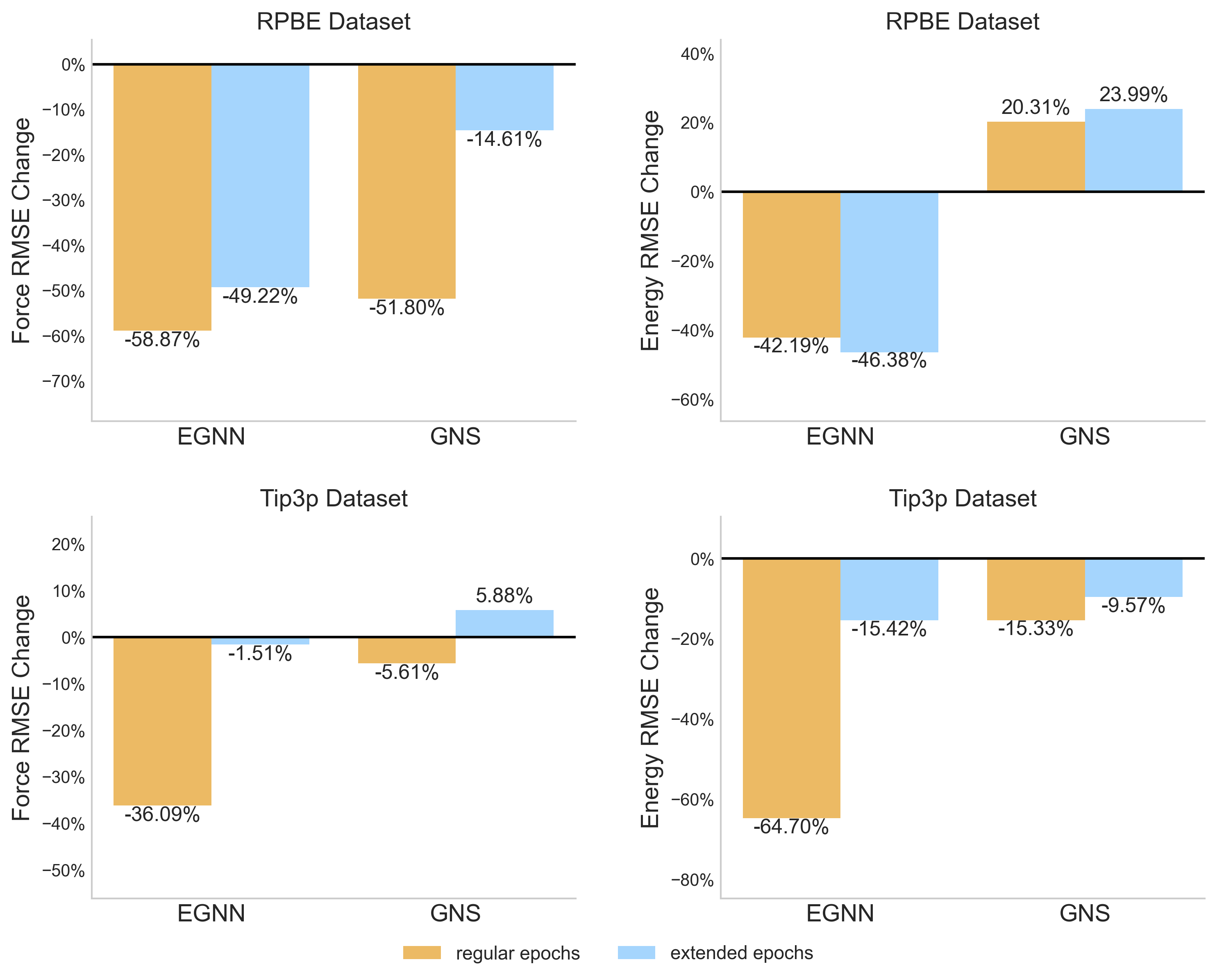
- Reduces Force RMSE by 47.48% and Energy RMSE by 53.45% for EGNN on the RPBE water dataset compared to training from scratch.
- Achieves consistent improvements across both equivariant (EGNN) and non-equivariant (GNS) architectures, showing model-agnostic benefits.
- Outperforms denoising pretraining on larger datasets (Tip3p), where denoising causes instability and degradation (e.g., 10x worse error for EGNN).
Breakthrough Assessment
7/10
Offers a robust, model-agnostic pretraining strategy that significantly improves data efficiency and stability for neural potentials, addressing limitations of the popular denoising approach in complex systems.