📝 Paper Summary
Masked Diffusion Models (MDMs)
Discrete Diffusion
Language Modeling
Biological Sequence Design
Path Planning (P2) enhances masked diffusion model inference by decomposing generation into planning and denoising steps, allowing the model to revisit and correct previously unmasked tokens.
Core Problem
Standard Masked Diffusion Models (MDMs) use a fixed, uniform unmasking order during inference, preventing the correction of mistakes made in earlier steps.
Why it matters:
- Uniform unmasking assumes a perfect denoiser, but real-world trained denoisers are imperfect, making random unmasking suboptimal.
- In domains like biological sequence design or reasoning, early errors propagate and cannot be fixed, degrading final sample quality.
- Current inference methods fail to unlock the full potential of MDMs, lagging behind autoregressive models in tasks requiring complex dependencies.
Concrete Example:
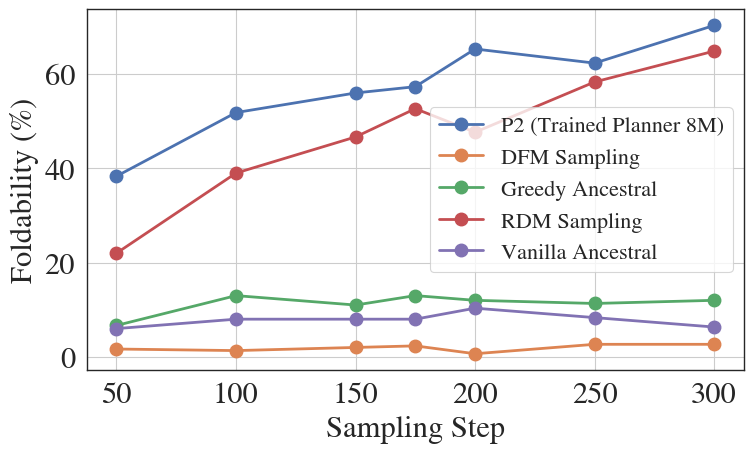
In protein generation, if an MDM incorrectly unmasks a residue early in the process that clashes with later structural constraints, standard inference keeps this error fixed, ruining the protein's foldability. P2 can identify this low-confidence token later and re-mask it for correction.
Key Novelty
Path Planning (P2) for MDM Inference
- Decomposes each generation step into two stages: a 'planner' selects which tokens to update (unmask or re-mask), and a 'denoiser' samples values for those tokens.
- Introduces a mechanism to 're-mask' and resample previously generated tokens that the model is least confident about, enabling self-correction during generation.
- Derives a new, expanded Evidence Lower Bound (ELBO) that theoretically justifies non-uniform, planner-guided generation trajectories.
Architecture
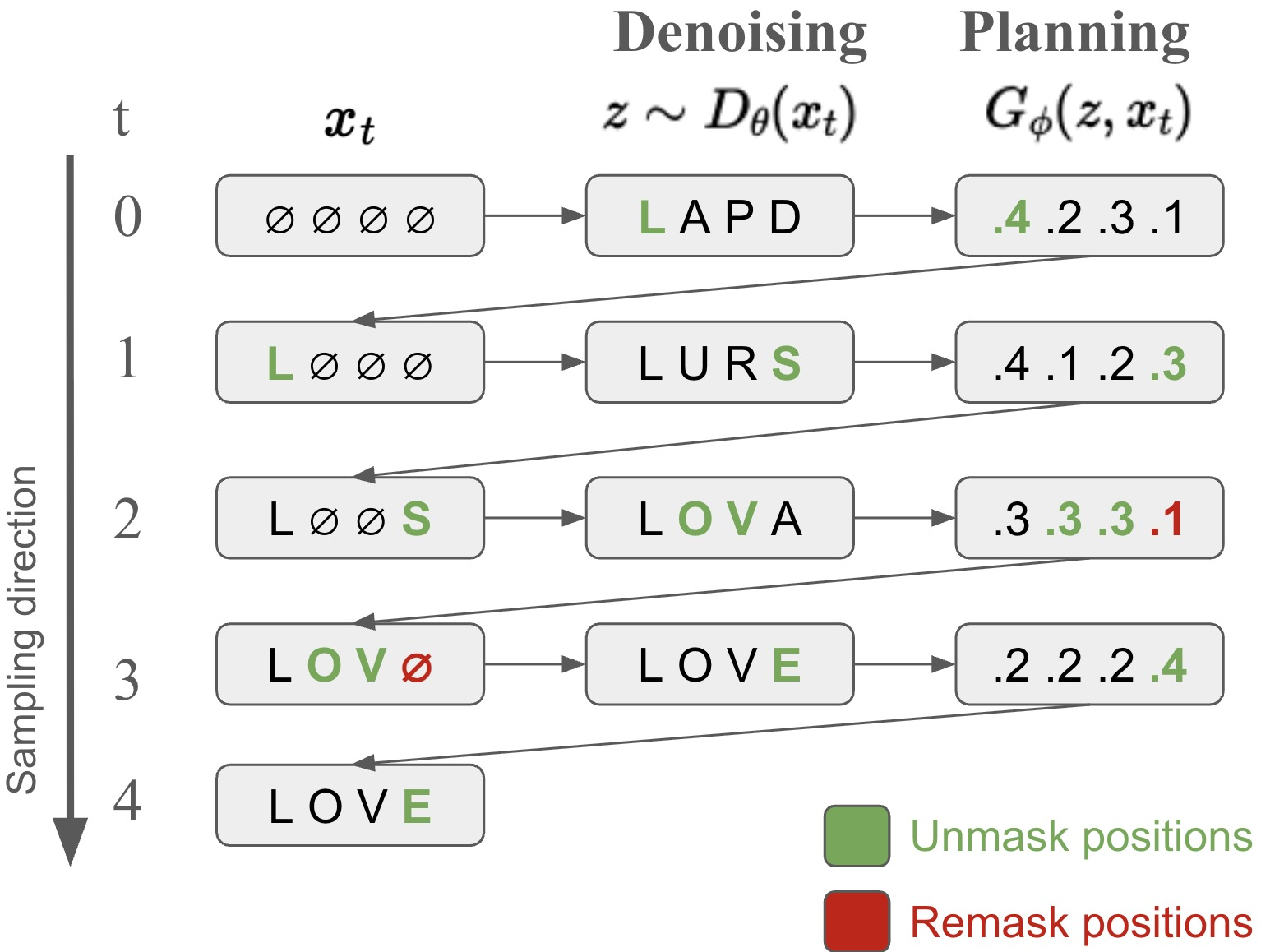
Conceptual diagram of the P2 inference process compared to standard MDM.
Evaluation Highlights
- +68% relative improvement in ROUGE score for story generation compared to standard MDM inference.
- +33% relative improvement in Pass@1 for code generation using a 1B parameter model, outpacing larger autoregressive baselines.
- +22% relative improvement in protein sequence foldability and +8% in RNA sequence pLDDT compared to state-of-the-art biological diffusion models.
Breakthrough Assessment
8/10
Significantly advances discrete diffusion by solving the 'fixed trajectory' limitation. theoretical grounding via the expanded ELBO and strong empirical gains across diverse domains (text, code, bio) suggest high impact.